Petersen Lab
Lab Rotation III
Division of Microbial Ecology, University of Vienna, Vienna, Austria
June 20, 2017 - July 21, 2017
Advisor: Julia Polzin, PhD
Project/Question I: Are sulfur oxidizing (SOX) symbiotic bacteria living in the feet of Lucinid clams?
Recently, sulfur binding proteins coded by SOX bacterial endosymbionts in lucinid clams have been detected in the feet of the clams. The feet search for sulfidic compounds in the sediment. As the bacteria need sulfide for their metabolism, the occurrence of these proteins in the feet are not surprising. However, these symbionts have only been thought to be located in the gills of the clams due to gas exchange (CO2, O2, etc) availability. Thus, it is hypothesized that the sulfur binding proteins are transported from these symbionts to the feet, rather than symbionts living in the feet, as well. No one has confirmed the absence of these symbionts in the feet. The aim of my project was to determine whether SOX symbionts lived in the feet of the clams, using FISH. The protocols I used did not detect bacteria at all in the feet of the clams. However, it is possible that due to autofluorescence and possible low abundance of bacteria, they were simply not detected. Use of other dyes and higher resolution microscopy could reveal the presence of bacteria, even if it's not the SOX symbionts. The DAPI stain (showing all DNA) of the feet looked super cool, even though no bacterial cells were seen. It's likely that the host cell DNA drowned out the signal of bacterial DNA.
Project/Question II: Where are Endozoicomonas bacteria in the gills of Lucinid clams?
Bacteria of the genus Endozoicomonas are common members of many animals' microbiomes, including coral, sponges, and fish. In some clams, bacteria of this genus are pathogenic, in others, they are harmless. Recently, Endozoicomonas bacteria have been found in the gill microbiome of Lucinid clams collected from Mauritania. This project aimed to determine where these bacteria are living in the gill, using FISH. Clams collected from Mauritania and Elba were both examined. The probe we used for Endozoicomonas contained the fluor dye. Sadly, fluor was known to exhibit high levels of autofluorescence for all probes used, including the SOX symbiont and EUB I-III probes. Thankfully, we had probes in other dyes for the SOX bacteria and EUBI-III. However, we only had Endozoicomonas probes in fluor. My lab rotation was not long enough for Endozoicomonas probes to be ordered in other dyes (like Cy5) to better search for the bacteria. But I'm looking forward to hearing their results when the new probes arrive.
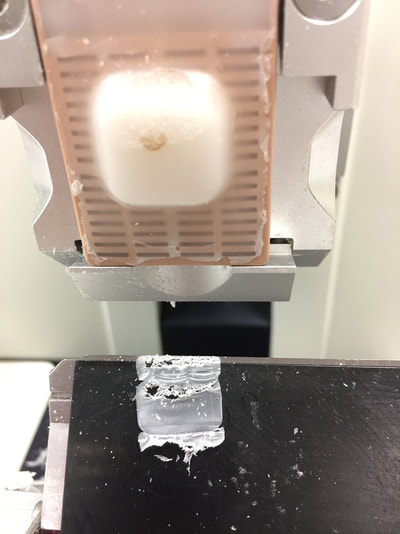
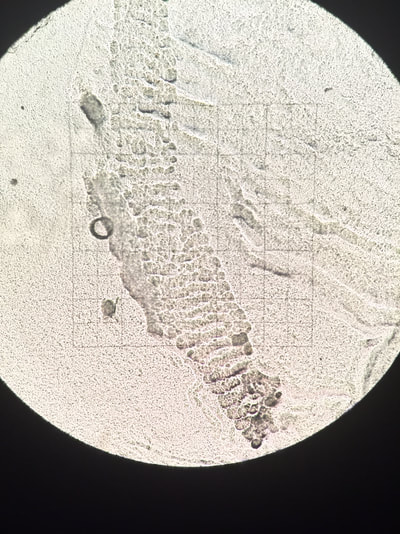
Below are some pictures from my project. :)
Lab Rotation III
Division of Microbial Ecology, University of Vienna, Vienna, Austria
June 20, 2017 - July 21, 2017
Advisor: Julia Polzin, PhD
Project/Question I: Are sulfur oxidizing (SOX) symbiotic bacteria living in the feet of Lucinid clams?
Recently, sulfur binding proteins coded by SOX bacterial endosymbionts in lucinid clams have been detected in the feet of the clams. The feet search for sulfidic compounds in the sediment. As the bacteria need sulfide for their metabolism, the occurrence of these proteins in the feet are not surprising. However, these symbionts have only been thought to be located in the gills of the clams due to gas exchange (CO2, O2, etc) availability. Thus, it is hypothesized that the sulfur binding proteins are transported from these symbionts to the feet, rather than symbionts living in the feet, as well. No one has confirmed the absence of these symbionts in the feet. The aim of my project was to determine whether SOX symbionts lived in the feet of the clams, using FISH. The protocols I used did not detect bacteria at all in the feet of the clams. However, it is possible that due to autofluorescence and possible low abundance of bacteria, they were simply not detected. Use of other dyes and higher resolution microscopy could reveal the presence of bacteria, even if it's not the SOX symbionts. The DAPI stain (showing all DNA) of the feet looked super cool, even though no bacterial cells were seen. It's likely that the host cell DNA drowned out the signal of bacterial DNA.
Project/Question II: Where are Endozoicomonas bacteria in the gills of Lucinid clams?
Bacteria of the genus Endozoicomonas are common members of many animals' microbiomes, including coral, sponges, and fish. In some clams, bacteria of this genus are pathogenic, in others, they are harmless. Recently, Endozoicomonas bacteria have been found in the gill microbiome of Lucinid clams collected from Mauritania. This project aimed to determine where these bacteria are living in the gill, using FISH. Clams collected from Mauritania and Elba were both examined. The probe we used for Endozoicomonas contained the fluor dye. Sadly, fluor was known to exhibit high levels of autofluorescence for all probes used, including the SOX symbiont and EUB I-III probes. Thankfully, we had probes in other dyes for the SOX bacteria and EUBI-III. However, we only had Endozoicomonas probes in fluor. My lab rotation was not long enough for Endozoicomonas probes to be ordered in other dyes (like Cy5) to better search for the bacteria. But I'm looking forward to hearing their results when the new probes arrive.
Below are some pictures from my project. :)